Evert-Jan Blom of Genetic Affairs developed a new AutoCluster Endogamy tool on GEDmatch together with Jarrett Ross of GeneaVlogger. Introducing it as AutoCluster Endo (AutoCluster Endogamy when you see it on GEDmatch) is a modified version of the AutoCluster clustering tool designed specifically for those dealing with endogamous matches. It was created to address analyzing endogamous matches more efficiently by filtering for the most relevant (shared) matches.
Thanks to Jarrett Ross for bringing up specific features he mentions in his video. It allows you to filter your primary matches by adjusting the average segment size, minimum largest segments, and number of largest segments. It also allows you to filter by your shared matches using the same filters as for primary matches and in addition, the total amount of shared cM between shared matches.
When I used to run the AutoCluster tool at MyHeritage, I noticed people would post their examples mentioning how endogamous their matches were or how burdensome, and problematic it was to deal with it. I also noticed a marked difference between their clusters and my own. For one, they had more than one cluster. I initially only had a single cluster until I uploaded one of my 1C1R with whom I do not share as much DNA (as expected I guess for someone of that relationship) and was enough for this tool to pick up. This cousin of mine appeared in my second cluster with other relatives on my paternal (non-Polynesian) side and he also produced gray squares matching several matches in the first/large cluster.
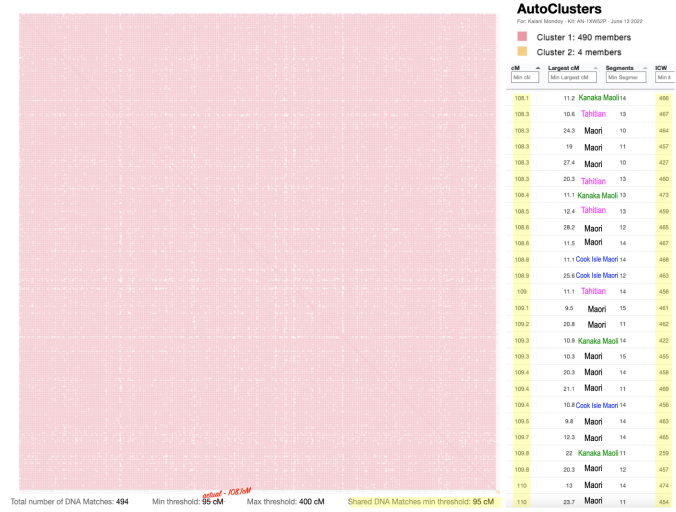
I emphasized in my AutoCluster for others to take note that the minimum threshold implemented was not 20cM or 30cM like many others that I remember seeing. Mine was significantly higher.
I also sorted my match list showing the lowest amount at the top, sharing 108.1cM, so the 26 matches I decided to show only shows from 108cM to 110cM. Of course there are 470 other matches that comprise that large cluster.
I kept pointing this out to others, how our minimum threshold will vary across different populations, depending on the amount of shared DNA we have our matches and the number of matches, etc. There is a bit more freedom with utilizing Genetic Affairs directly.
With this AutoCluster Endogamy tool at GEDmatch, you can do quite a bit. This tool is offered to Tier 1 subscribers (Tier 1 pay-as-you-go membership $15 per month and Recurring monthly Tier 1 memberships $10 per month) only.
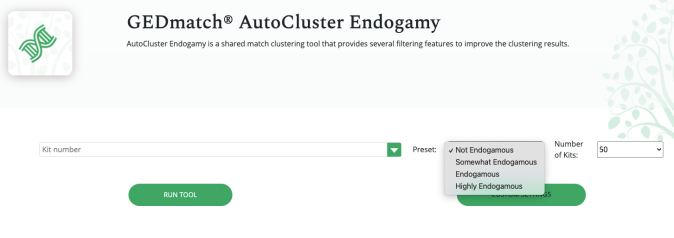
The first thing you will notice is that you have the option to select the level of your endogamy or how endogamous you are.
The default is set to “Not Endogamous.” While I only tried the “Endogamous” option to see the difference from the “Highly Endogamous” (Polynesians should be using “Highly Endogamous”) and noticed that the parameters were set higher to numbers that are very familiar to me.
Leah Larkin (The DNA Geek) has shown in her presentations charts of various endogamous populations and to what degree of endogamy each has to deal with. This is where I first saw how she utilized the average size segment to quantify endogamy, how to gauge how much endogamy you are really dealing with.
She took the amount of shared DNA for Close Relatives (Ancestry), predicted First Cousins, Second Cousins, Third Cousins, Fourth Cousins and Distant Cousins, divided by the number of segments to come up with the average size segment. What was presented were various sizes present in specific endogamous populations. She had mild, moderate and strong endogamy. These were average size segments present in specific (predicted) relationships, i.e. 1C, 2C, 3C, etc.
In her comparison, the one that had the smallest average size segment were Polynesians. She also separated to demonstrate what Western Polynesians had compared to Eastern Polynesians. She has confirmed (although many of us probably noticed this already) how endogamous, or extremely (“Highly” is the term used for this AutoCluster Endogamous tool at GEDmatch) endogamous Polynesians are. This could not be done without the help of others submitting their samples to Leah for analysis. I was able to submit one Samoan and two Kanaka Maoli samples to her to utilize. And the results were worth it!
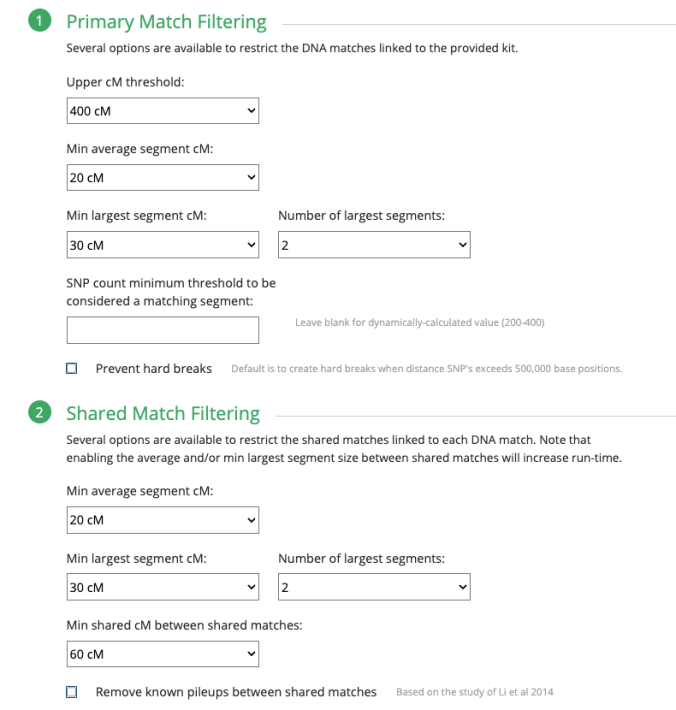
Having said all of that, do know that Polynesians should automatically select “Highly Endogamous.” This seems to raise the Min average segment cM and other parameters. This image below is an example of what it looks like when you do not select anything and keep it at the default “Not Endogamous.”
Even with “Not Endogamous” you can still adjust the settings to your liking.
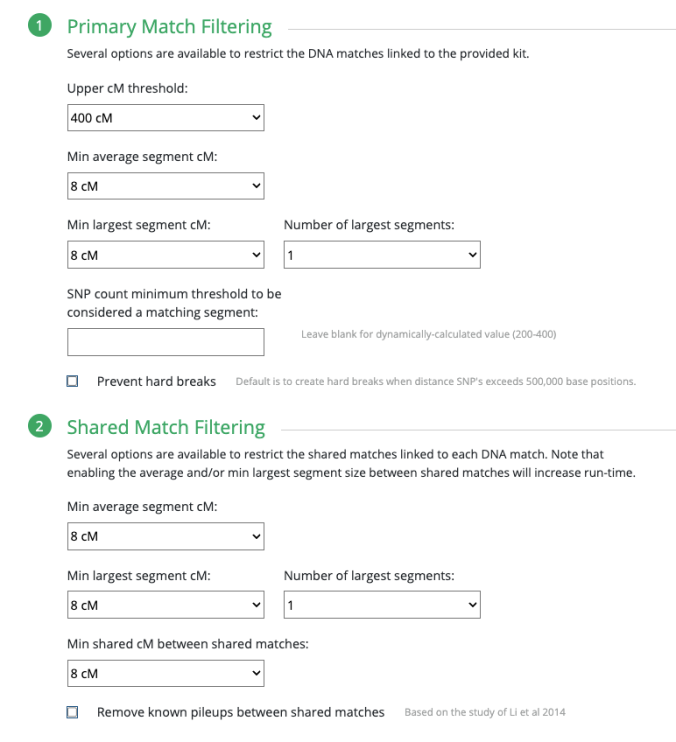
So below are the settings that you would automatically see when selecting “Highly Endogamous.”
It is important to note, based on what I have seen others post with their own comparison and my 11 years of noticing the largest segment size among Polynesians and known relationships, that the Min largest segment cM selected for 30cM is a good minimum amount to use. This is what you would expect around the 2nd Cousin level.
I have at Ancestry and MyHeritage (as do other relatives of mine) endogamous matches whose largest segment exceeds 30cM yet what helped distinguish it from a true close relative versus an endogamous one is how they still have a significantly high amount of segments.
Below is a table of all of my matches (Ancestry) and I have highlighted my known relatives. The ones not highlighted are the endogamous matches.
You can clearly see how with my known (highlighted) 2C, 2C1R, and 3C1R relatives (Predited as Second Cousin) the number of segments aren’t always as high. The ones that are, they have little to no non-Polynesian lines, which means more Hawaiian branches that are coming up as matches to me. But, the largest segment is coming from our most recent common ancestor. Notice that for the New Zealand Maori and Kanaka Maoli matches the number of segments are really high.
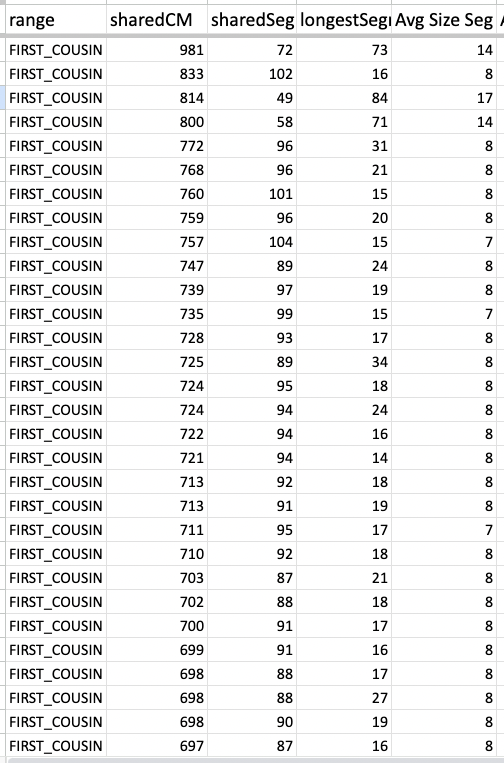
For comparison, this (table below) is a cousin of mine. Although I did not indicate the true close relatives, it should be obvious based on the high amount of segments plus the average segment size which ones are truly close relatives.
For the past 11 years, this is what I have been noticing. That it was not common to see DNA matches among Polynesians (mainly Kanaka Maoli and NZ Maori) whose largest segment size exceeded 20cM. Utilizing the average size (taking the total shared cM divided by the number of segments), we see 7cM and 8cM to be the norm both in my cousin’s predicted First Cousin matches and my predicted Second Cousin matches. It is pretty common even when looking at the 3rd Cousin, 4th Cousin, and Distant cousin matches.
So this is why we have the type of results you would see with autoclustering and why the need to be able to adjust these parameters in order to find the best matches (true close relatives) to be used in clustering.
So now we have an understanding of what to expect among Polynesian DNA matches as far as the average size segment, the number of segments (to help get the average size segment), and the largest segment size. In my next blog entry, I will address the results of running this tool and how adjusting these may or may not be as useful.
One thing to note is that various companies will use the longest block (FTDNA), longest segment (Ancestry), and largest segment (MyHeritage & GEDmatch) for the same thing. I may use these terms interchangeably, but for this particular GEDmatch tool, I’ll only refer to it as largest segment.
This is so helpful! Thank you Kalani!
LikeLike
You’re welcome! Still playing around with this tool, seems like there is some potential.
LikeLike
Pingback: AutoCluster Endogamy tool at GEDmatch.com (Part 2) | Polynesian DNA
Thanks for the blog Kalani! Nice case study.
LikeLike